A group from Marine Biotechnology Research Center, Korea, etc. has reported on inhibition of SARS-CoV-2 infection with seaweed polysaccharides and those cytotoxicity.
https://www.ncbi.nlm.nih.gov/pmc/articles/PMC8071526/
Seaweeds are an excellent source of bioactive compounds such as polysaccharides, dietary fibers, amino acids, essential fatty acids, carotenoids, phlorotannins, vitamins, and minerals. These compounds have been reported to have a variety of pharmacological activities such as antitumor, antiviral, antioxidant, antimicrobial, anticoagulant, and immune-inflammatory effects
Authers investigated the inhibitory activities of the following crude polysaccharides (CPs):
Undaria pinnatifida sporophyll (CPUP),
Laminaria japonica (CPLJ),
Hizikia fusiforme (CPHF),
Sargassum horneri (CPSH),
Abalone viscera (CPAV),
Codium fragile (CPCF),
Porphyra tenera (CPPT), and
Fucoidan.
Cytotoxicity Assay with the viability of HEK293/ACE2 cells was assessed. When the HEK293/ACE2 cells were treated with each CP at the final concentration range from 1 ng/mL to 1 mg/mL (serial diluted 1/10) for 96 h, most of the CPs did not show severe cytotoxicity. However, CPAV, CPCF, and CPPT were slightly cytotoxic at a concentration of 1 mg/mL. CPAV showed notable cytotoxicity when applied at a concentration of 1 mg/mL, following a degree of cytotoxicity by CPPT, CPCF, CPHF, CPLJ, CPUP, CPSH, and fucoidan. Nevertheless, the CC50 of all CPs was over 500 μg/mL.
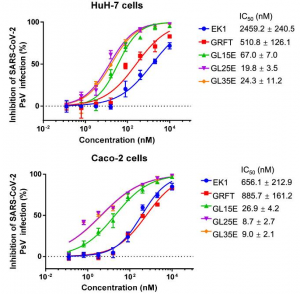
The testing of the inhibition of viral infection by CPs was performed with a SARS-CoV-2 pseudovirus. All CPs, except CPPT, inhibited SARS-CoV-2 pseudovirus infection of HEK293/ACE2 cells at various concentrations. Among the CPs tested, CPSH showed the strongest antiviral activity with an IC50 of 12 μg/mL, followed by CPAV (33 μg/mL), CPHF (47 μg/mL), CPCF (74 μg/mL), CPLJ (105 μg/mL), fucoidan (142 μg/mL), and CPUP (289 μg/mL), respectively.
As a conclusion, Sargassum horneri (CPSH) would be the best as an anti-SARS-CoV-2 agent.