Glyco26 was held at Academia Sinica in Taipei from August 27 to September 1, 2023.
First of all, I would like to summarize the trends in lectin arrays (also called lectin microarrays).
As you may know, lectin arrays appeared in 2005, and two groups, Lara Mahal from the United States and Jun Hirabayashi from Japan, published papers on glycan profiling analysis technology using lectin arrays. The following is one scene of Lara Mahal’s presentation (D010) from the University of Alberta at Glyco26.

Reference:A group of Lara Mahal et al.
Reference:A group of Jun Hirabayashi and Atsushi Kuno et al.
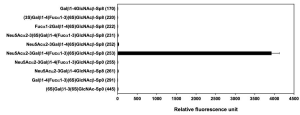
The latter paper adopted an evanescent-field fluorescence excitation method as the excitation technology to get a precise glycan profiling using a lectin array. This method enables detection of very weak biomolecular interactions between lectins immobilized on slide glasses and fluorescently labeled glycoproteins as analytes, non-destructively and directly from the liquid phase. And further, the admin. of this blog was also a co-author of this paper, and this technology was placed on the market as a glycan profiler (named GlycoStation® Reader 1200) in 2007 by myself.
Regarding the progress of researches using lectin arrays or on the lectin arrays themselves in Glyco26, regarding the former, there were presentations from a group of Academia Sinica and a group of AIST.
Wu-Show Su et al., Academia Sinica used lectin arrays for glycan profiling of IL6 secreted from cancer cells (A107), and Atsushi Kuno et al., AIST were verifying the differences in glycosylation at each local tissue site of cardic fibrosis using paraffin-embedded samples. In this study, an advanced glycan profiler, GSR2300 made by emukk LLC, was used, and it was shown that it is possible to get glycan profiling from only 3 cells (C050).
Regarding lectin arrays themselves, two groups gave presentations on arraying human lectins.
Kurt Drickamer et al., ICL develped an array immobilized with a total of 31 types of human lectins, including C-type lectins, Galectins, Siglec, and so forth to screen the binding specificity of these human lectins to pathogenic and commensal micro-organisms. This microarray provides unique information on how various lectins, which are at the forefront of human innate immunity, react with various microorganisms, and this demonstrates the usefulness of human lectin arrays (D026 ). In this study, human lectins were biotinylated and those were immobilized on a streptavidin-covered substrate. In addition, GlycoGenetics announced a microarray product that immobilizes 11 types of human lectins, mainly galectin (C072). Many of the existing lectin arrays are using plant lectins, such as LecChip, and lectin arrays adopting human lectins are a new trend in lectin arrays. However, the glycan binding specificity of human lectins is mainly related to the innate immune system, and therefore the coverage of glycans is not good enough. From the viewpoint of glycan profiling, existing lectin arrays centered on plant lectins are sufficiently powerful, and further they are highly comprehensive (for example, you can refer to arrays using plant lectins (C053) by Jaroslav Katrilik et al. of Slovak Academy of Sciences). So, existing lectin arrays and human lectin arrays will habitat segregation according to the applications, I think.
Next, regarding glycan arrays, as you know, the US CFG glycan arrays have become the de-facto position due to their high comprehensiveness. There were several presentations using this glycan array to evaluate the glycan binding specificity of glycan-binding proteins. One of the most interesting presentation at Glyco26 using glycan arrays would be “Blood Group A enhances SARS-CoV-2 infection” presented by Shang Chuen Wu, Richard Cummings and Sean Stowell et al., Harvard Medical School (A071). Ryuhei Hayashi and Yashuhiro Ozeki et al., Yokohama City Univ. used a unique glycan array (manufactured by GlycoTechnica) to evaluate the glycan binding specificity of marine sponge’s lectins . A newly discovered lectin named Chal18 was shown to have very strong specificity for T-antigen and to exhibited strong cytotoxicity against colorectal cancer cells (A031).
A new trend regarding glycan arrays is the movement to create arrays of glycans using pathogenic bacteria. Laurriel Macali and Todd Lowary et al., Academia Sinica was discussing the glycan binding specificity of mycobacteriophages using glycan arrays that immobilize 66 glycans exstracted from microorganisms. In this glycan array, glycans were immobilized on a slide glass using the neoglycoprotein method (i.e., a method in which a conjugate of an extracted glycan and BSA is made and the BSA is immobilized on the slide glass) (A040).
Microorganisms have a wide variety of glycans, and also their purification is difficult. Furthermore, since the state of glycans actually expressed on the surface of microorganisms and the state of extracted glycans and immobilized on glass slides are not the same, Hau-Ming Jan and Sean Stowell et al., Harvard Medical School proposed a method named MMA (microbe microarray) in which the microorganisms are directly arrayed on a slide glass, and through studies on interaction of Gal-8 with glycan arrays and MMAs as an example, they concluded that MMA could predict binding properties more accurately than conventional glycan arrays (A048).
At Glyco26, there were a total of 343 presentations, so the admin. might not be able to cover them all and might have overlooked some. I appreciate your understanding about it and I hope this information is of some help to you.