A group from Department of Physics, University of California, San Diego, etc. has reported about prediction of glycan structures from lectin binding yusing a Boltzmann model prediction.
https://www.ncbi.nlm.nih.gov/pmc/articles/PMC10274649/
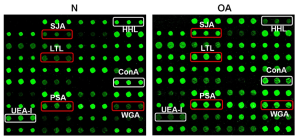
In this study, it was tested if glycan structures can be predicted from lectin binding patterns. Each lectin has a small set of motifs to which it binds. Since lectins are not expected to have logically complex binding rules, it was thought that shallow neural network topologies to be adequate. A reasonable minimal model for this interaction would be hought to be a fully visible Boltzmann machine, which can be conceptualized as a two layer network.
To be honest, I think that it’s not so accurate than expected. This will be partly because the glycans are a very heterogeneous group, and partly because the binding specificity of lectins is ambiguous. There are many people who are against glycan structure estimation using this kind of lectin panels, and there are a large number of MS/MS adherents.
However, in the context of this paper’s discussion, their thought is the same as mine.
“Information got from lectins is accurate enough in a biological context!”