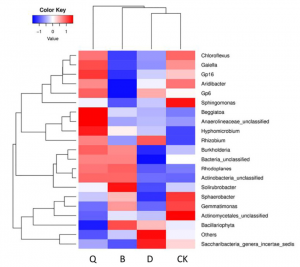
A group from Institute of Microbiology, Chinese Academy of Sciences, Beijing, China, has studies bacterial and fungal communities of chili pepper rhizosphere using amplicons (16S and ITS) and investigated how Fusarium wilt disease (FWD) affects its microbiomes.
https://www.ncbi.nlm.nih.gov/pmc/articles/PMC8444440/
Fusarium wilt disease (FWD) is often caused by the Fusarium oxysporum species complex, a classical soil-borne disease that attacks a wide variety of economically important crops, including banana, watermelon, and Solanaceae plants (e.g., tomato, eggplant, and chili pepper).
The relative abundance of several potential pathogenic fungi from the genera Diaporthe, Fusarium, Phomopsis, Plectosphaerella, Stemphylium, and Cryptococcus was also significantly higher in the diseased plant root, and several potential beneficial bacteria from the genera Pseudomonas, Streptomyces, Klebsiella, Enterobacter, Microbacterium, Bacillus, Chitinophaga, and Citrobacter were significantly enriched in the diseased plants.
Several functional genes involved in plant-microbiome signaling pathways were more abundant in the microbiome of the diseased root endosphere than in the healthy. For instance, the relative abundance of genes associated with methyl-accepting chemotaxis proteins (MCPs) was increased by 33.2–218.2% in the microbiome of the diseased root endosphere, compared with the healthy plant. The relative abundance of the functional genes associated with the downstream of MCPs, such as histidine kinase CheA and purine-binding chemotaxis protein CheW, also increased by 15.0–40.3% in the microbiome of the diseased root endosphere, compared with the healthy plant.
Several genes encoding MCPs associated with plant-microbiome signaling pathways were enriched in the microbiome of diseased root endosphere. MCPs are the predominant chemoreceptors in motile bacteria that alter the activity of CheA histidine kinase and the bacterial swimming behavior upon detection of specific chemicals . MCPs have been identified in typically beneficial bacteria, e.g., Bacillus subtilis and Pseudomonas spp. , which were also significantly enriched in diseased plant in the current study. Under stress conditions, such as pathogen invasion, a plant can attract distant beneficial microbes by actively releasing nonvolatile root exudates, such as amino acids, nucleotides, and long-chain organic acids, or by actively emitting blends of volatile organic compounds. The findings of the current study suggest that the MCP gene enrichment in diseased plants may be related to the response of MCP-producing bacteria to plant-released signal molecules. These bacteria would use MCPs to detect specific concentrations of these signaling molecules in the extracellular matrix, enabling directional accumulation of the bacteria to the plant. Further research will be required.
This study provides evidence on the critical role of bacterial taxa in the “cry for help” strategy of the host plant, in which the plant actively involves its microbial partners to maximize its or its offspring survival and growth under external stress.