A group from the collaborative network of the SARS-CoV-2 Assessment of Viral Evolution (SAVE) program of the National Institute of Allergy and Infectious Diseases (NIAID) has reported that SARS-CoV-2 omicron variant shows attenuated lung disease in mice and hamsters, which parallels preliminary human clinical data.
https://www.ncbi.nlm.nih.gov/pmc/articles/PMC8722607/
Some recent studies (as also introduced in this blog) speculated that because mutations in the omicron spike protein overlap with ones known to promote adaptation to mouse hosts, the progenitor of omicron jumped from humans to mice, and then back into humans. In support of this, omicron but not Wuhan-1 RBD binds to murine ACE2. Hamsters also are highly susceptible to SARS-CoV-2 infection with similar pathological changes seen in lung tissues from COVID-19 patients. Using experimental data from multiple laboratories of the SAVE/NIAID consortium, authors report on the infectivity and pathogenesis of multiple omicron isolates in mice and hamsters.
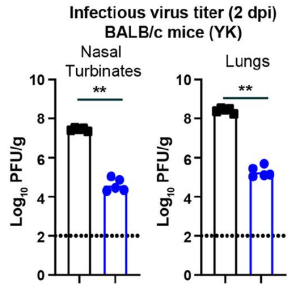
Here is the data of comparison between omicron in BALB/c mice and beta variant. At 2 days-post-infection (dpi), infectious virus levels in the nasal turbinates and lungs were significantly lower (~1,000-fold, P < 0.001) in BALB/c mice infected with omicron compared to beta virus.
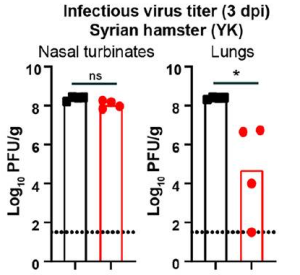
A comparison of infectious viral burden in hamster tissues at 3 dpi between delta and omicron viruses showed virtually no difference in nasal turbinates but substantially less infection of omicron in the lungs of hamsters.
Here is the comparison of lung pathology in Syrian hamsters after infection with delta or omicron viruses. Macroscopically, the lungs obtained from the delta-infected hamsters showed congestion and/or hemorrhage, but this was absent in omicron-infected animals. Overall, the Omicron variant replicates less efficiently in the lungs of Syrian hamsters, which results in less severe pneumonia compared to the delta variant.
Our experiments suggest that compared to other SARS-CoV-2 isolates (e.g., beta or delta), the Omicron variant infection is attenuated in laboratory mice and hamsters for causing infection and/or disease. While these results are consistent with the very preliminary clinical data in humans suggesting that omicron causes a more transmissible yet possibly milder respiratory infection, the basis for the attenuation in rodents remains unknown.
One pre-print study (also introduced in this blog) suggests that omicron9 replicates faster in the human bronchus and less in lung cells, which may explain its greater transmissibility and putative lower disease severity; although it remains unclear if these observations extend to rodents, we observed less infection of hamster bronchial cells in vivo with omicron than delta virus. Lower viral burden in nasal washes and turbinates in 129 mice were also measured comparing to other SARS-CoV-2 strains. The attenuation in mice was unexpected given that omicron has multiple mutations in the RBD that are sites associated with adaptation for mice.
Whereas the more than 30 substitutions in the omicron could impact receptor engagement and cell entry, sequence changes in other domains could affects replication, and so forth. Thus, detailed genetic and functional studies are required to define the basis of virological and clinical attenuation of omicron in mice and hamsters.